BrainGEP - In silico gene prioritisation with brain gene expression data
BrainGEP (Brain Gene Expression Prioritisation) is an R package which performs in silico gene prioritisation using the Allen Human Brain Atlas developing and adult human brain microarray data.
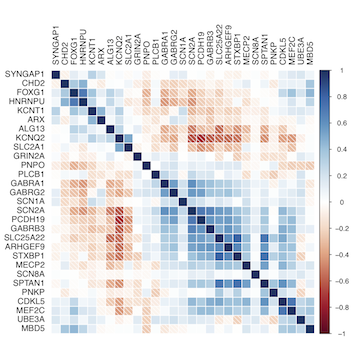
The package currently uses Pearson's and Spearman's correlation coefficients to assess the correlation between a set of known disease causing genes and prioritises candidate genes with respect to the known disease genes. The analysis can be done across the whole brain for both adult and developing brains, as well as for cortex (CX), cerebellum (CB) and brainstem (BS) regions across the adult human brains, and cortex (CX) and other (non-CX) areas across the developing human brains. The package requires separate downloading of the Allen Human Brain Atlas data.
In the UNIX operating system, the BrainGEP tarball can be installed using the following command:
R CMD INSTALL -l /home/your_username/R_libs BrainGEP_1.0.4.tar.gz
The user can then go into R and type
require(BrainGEP,lib.loc="/home/your_username/R_libs")
The package is now ready for use.
The data initially needs to be processed to get it into a form suitable for our analysis. It also requires the user to specify a path where the processed data is to be saved (save_path) and then re-loaded (using the Load_AHBA_data function) for each subsequent use of the package.
Please refer to Instructions to see how to begin using the package and the BrainGEP manual for detailed instructions in using the functions of the package. Example R code showing how to use the functions is available here.
An example application is published in Oliver, Lukic et al in which 179 candidate Epileptic Encephalopathy (EE) genes are prioritised with respect to 29 known EE genes. Please cite this paper if you use BrainGEP.
The adult AHBA microarray data has been slightly updated in September 2013. A document showing the very minor effects of these changes is available here.
This package is written and maintained by Vesna Lukic. Last modified 25th August 2015.